I recently upgraded to Python 3.10, and because of some problems with my previous installation I decided to do a complete new installation using pip, which required installation of all the libraries needed for my various programs.
For my linear algebra functions based on the Scipy library I have also installed PyPardiso which provides substantially better performance than the Scipy functions for solving large sparse systems. As reported here I previously found I had to search for a non-standard conda library for the installation, but this time the latest documentation recommended installation with pip: pypardiso 0.4.1
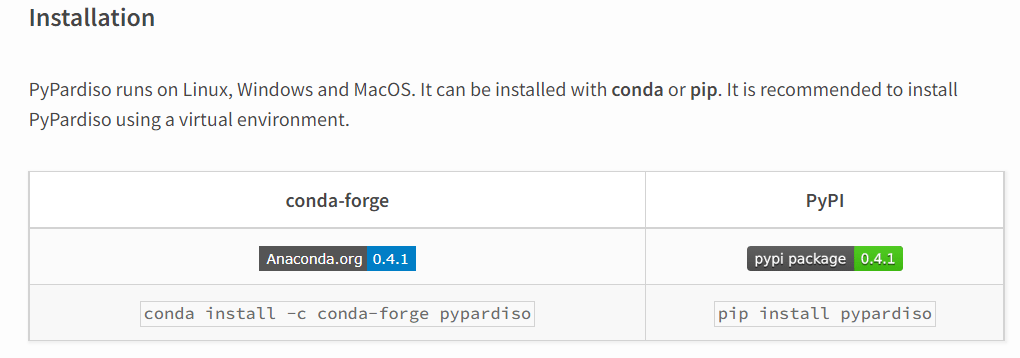
Since PyPardiso calls the Intel MKL library, I installed this first, so my installation procedure was:
- Install the MKL library: pip install mkl
- Install PyPardiso: pip install pypardiso
Code for using the PyPardiso spsparse function (or optionally the equivalent Scipy.sparse function) can be found at: Installing PyPardiso.
Checking the performance of the equivalent functions, the pyPardiso spsolve function was found to be much faster than the Scipy equivalent, as previously reported at: Making Finite Element Analysis go faster – Update and PyPardiso.
The Scipy sparse.spsolve function was found to have reverted to the slower times reported at: Making Finite Element Analysis go faster … The change in performance is (probably) related to installation of the scikit-umfpack library, which I had not installed at the time of the tests, but even after umfpack is properly installed, the pyPardiso function is much faster than any option with the Scipy function.

Pingback: Installing PyPardiso | Newton Excel Bach, not (just) an Excel Blog
Pingback: Speed of Scipy Linear Algebra Solvers | Newton Excel Bach, not (just) an Excel Blog
Pingback: Scipy Functions with Excel and pyxll 7 – Linear Algebra | Newton Excel Bach, not (just) an Excel Blog
Pingback: Scipy update and Linalgfuncs speed check | Newton Excel Bach, not (just) an Excel Blog