Today it was announced that Facebook, the meta company that owns Facebook, was going to be renamed “Meta”, which reminded me of an xkcd episode from 10 years ago:


Update 5th June 2022: The PyPardiso package may now be installed simply with pip (see Installing PyPardiso and speed of Scipy spsolve):
The PyPardiso package provides an interface to the Intel MKL Pardiso library to solve large sparse linear systems of equations. Trying to install this package with Conda raises the message:
To search for alternate channels that may provide the conda package you’re
looking for, navigate to https://anaconda.org and use the search bar at the top of the page.
This provides the link: https://anaconda.org/haasad/pypardiso
which provides the command line:
conda install -c haasad pypardiso
Ensure that the Intel MKL library is installed before installing pypardiso.
The pypardiso Github site is at:https://github.com/haasad/PyPardisoProject
This provides the following documentation:

I have now modified my py_spsolve function to use the PyPardiso solver by default:
@xl_func(disable_function_wizard_calc=True)
@xl_func(category = "Scipy-Linalg")
@xl_arg('v', 'numpy_array', ndim=1)
@xl_arg('i', 'numpy_array', ndim=1)
@xl_arg('j', 'numpy_array', ndim=1)
@xl_arg('ya', 'numpy_array', ndim=1)
@xl_return('numpy_array')
def py_spsolve(v, i, j, ya, out = 0, use_pardiso = True, permc_spec = 'MMD_AT_PLUS_A'):
"""
Solve a matrix equation Ax = y in sparse COO format.
:param v: value vector
:param i: row index vector
:param j: column index vector
:param ya: y vector
"""
timea = np.zeros((1,2))
stime =time.perf_counter()
n = ya.size
A = ssp.coo_matrix((v,(i,j)),shape=(n,n)).tocsc()
timea[0,0] = time.perf_counter()-stime
if use_pardiso:
res = spsolve(A, ya)
else:
res = sspla.spsolve(A, ya, permc_spec)
timea[0,1] = time.perf_counter()-stime
if out == 1:
return timea
else:
return res
The Chieftains are a traditional Irish folk band who have been performing for nearly 60 years. Paddy Moloney was one of their founding members, and died in hospital on 12th October. He has received widespread tributes, including from the President of Ireland, Michael D. Higgins:
“The Irish music community, and indeed the much larger community throughout the world who found such inspiration in his work, will have learned with great sadness today of the passing of Paddy Moloney. […] Paddy, with his extraordinary skills as an instrumentalist, notably the uilleann pipes and bodhrán, was at the forefront of the renaissance of interest in Irish music, bringing a greater appreciation of Irish music and culture internationally.”
In my opinion some of their best work was produced with other World renowned artists, of which the performance below with Sinead O’Connor is an exceptional example:
For more on the history of the Chieftains see:
The previous post on this topic looked at the performance of alternative Scipy sparse equation solvers. This post updates those results with the current Scipy version (1.7.1), with very different results. It also compares the Scipy solver performance with the PyPardiso package, which provides an interface to the Intel MKL Pardiso library to solve large sparse linear systems of equations.
The sparse solver function spsolve has an optional argument called permc_spec that defines “how to permute the columns of the matrix for sparsity preservation”. My timings for the previous post found that the argument NATURAL had very much better performance than any other alternative. This time, NATURAL performed as before, but all the other options had substantially improved, so that it had become the second slowest for the matrix used in the previous post, and the slowest for larger matrices.
The solvers have been compared for solving the stiffness matrix for 3D building frames of increasing size. The table below shows the timing results for the four spsolve options, the banded solver, two iterative solver options, and the PyPardiso solver. Each option has been timed for a system with 14742 equations (as in the previous post), 29487 equations, and 56705 equations.
Two times are reported for the Pardiso function: the time for the first calculation, and a much faster time for subsequent calculations, provided the stiffness matrix is unchanged.
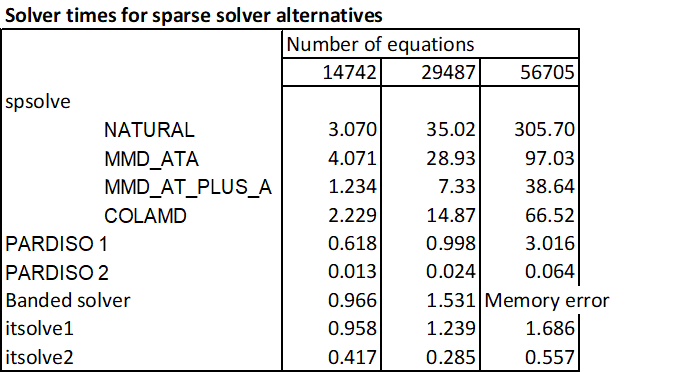
In summary the Pardiso package was substantially faster than any spsolve option, especially for very large systems, was also faster than the banded solver, and was able to handle larger systems than the banded solver.
In the next post I will look at procedures for installing the Pardiso package, and calling the function from Excel using pyxll.
Python 3.10 was released on October 4th, rapidly followed by a new release of pyxll (5.3.0), but the Anaconda site does not seem to have it yet, so is it time to upgrade or not. A search on that question found:
Python 3.10 is now available–but should you switch to it immediately? And if not now, when?
The short answer is, no, you probably don’t want to switch immediately; quite possibly you can’t switch immediately. To understand why, we need to consider Python packaging, the software development process, and take a look at the history of past releases.
We can then make a guess about when Python 3.10 will actually be usable.
The problems with a new major Python release
I’m writing this article on October 5th, 2021, the day after 3.10 was released… and far too early to start using Python 3.10.
As with many open source projects, Python, Python libraries, and most of the toolchain and packages mentioned in this article are based on volunteer labor. None of this should be read as a complaint towards the people doing the maintenance, they’re doing hugely valuable work for free, and everything takes time. In addition, because upgrades involve so many different groups, coordination and releases take even more time.
With that in mind, let’s consider the problems with using 3.10 today: …
by Itamar Turner-Trauring